This research uses spatial transcriptomics to map interactions between T cells, cancer cells, and immunosuppressive cells in tumours. Findings suggest cancer suppresses immune responses by surrounding and weakening T cells. The work aims to improve immunotherapy and enable personalised cancer treatment through detailed tumour mapping.
This thesis examines how octopuses respond to climate change at a molecular level, focusing on ocean acidification and RNA editing. Rising temperatures harm octopus reproduction, growth, and survival, while acidification produces mixed effects—some species show stress, yet others demonstrate resilience. Cephalopods overall appear more tolerant of acidification than fish, raising questions about the mechanisms behind this adaptability. Thousands of acidification-responsive edits disproportionately affect C2H2 zinc finger regulators, altering predicted binding targets, including nuclear pore components implicated in stress responses.
This research applies machine learning to genetic data to distinguish harmless DNA variations from cancer-causing mutations. By treating DNA like a crime scene, the model learns to identify which genetic changes truly drive breast cancer risk, supporting more accurate diagnosis and informed clinical decision-making.
This research investigates HMGN proteins, which organize the genome and help cells access the correct genes. By mapping their activity and removing them with CRISPR, the study shows that HMGNs act as DNA “librarians.” Their dysfunction leads to gene misregulation linked to many diseases.
About 8% of the human genome originates from ancient viruses. This research uses bioinformatics and evolutionary comparisons to understand why viral DNA persists and how cells silence it through DNA methylation. Identifying how genomes separate useful from non-functional DNA helps clarify which genetic elements matter for human health and disease.
This research introduces a computational method that detects up to one trillion RNA viruses hidden in standard RNA-sequencing data. By targeting protein signatures shared across all RNA viruses, the approach reveals viral RNA that previously went unnoticed. This enables large-scale viral discovery, tracking, and potential breakthroughs in understanding disease mechanisms.
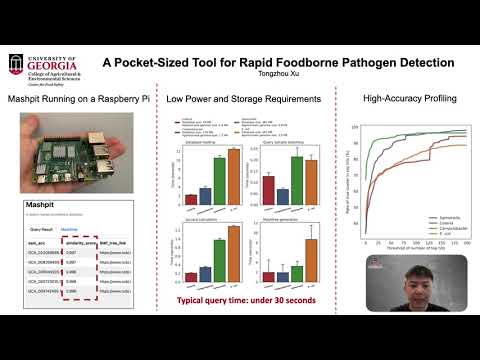
Mashpit is a portable genome-search tool that runs on a Raspberry Pi, enabling rapid, offline screening of Salmonella genomes. Using MinHash sketches, it scans hundreds of thousands of genomes in seconds, offering small or low-resource labs a fast, accessible way to identify related isolates before performing high-resolution follow-up analyses.