This talk traces the devastation of the Black Death to highlight a modern crisis: antibiotic resistance. Misuse of antibiotics accelerates the rise of superbugs. Using AI and machine learning, the research identifies genetic resistance patterns and guides effective treatments, aiming to improve clinical decisions and prevent a return to a pre-antibiotic era.
This research examines how microorganisms in maple sap influence the quality of maple syrup. By studying bacteria such as Pseudomonas and Duganella, the project explores how environmental factors like temperature and iron availability shape microbial interactions during the tapping season, ultimately affecting syrup flavor, color, and overall production.
This research investigates how a gonorrhea protein is processed in E. coli using cellular signal sequences, which act like "ZIP codes" directing the protein to its proper location. By identifying effective signal sequences, the study informs potential molecular targets for earlier detection and better treatment, aiming to prevent gonorrhea-related infertility and improve women's reproductive health.
This research investigates how bacteria develop resistance to antibiotics, a growing global health threat. By identifying resistant bacteria and analysing how they chemically modify antibiotics, the work aims to uncover resistance mechanisms. These insights are essential for preserving antibiotic effectiveness and safeguarding treatments against life-threatening infections.
This research tackles antibiotic resistance by developing nano-scale microfluidic cultures that isolate and study previously unculturable bacteria. By screening rare microbes and directly testing their antimicrobial activity, the platform accelerates discovery of new antibiotics, offering a powerful tool against drug-resistant superbugs.
This research investigates how the human microbiome protects against Streptococcus pneumoniae. Focusing on Streptococcus mitis, it shows how beneficial bacteria detect chemical signals from pathogens and block infection. Understanding when this microbial “security system” succeeds or fails may lead to new strategies for preventing disease.
This research explores how bacteria choose between free-swimming and biofilm lifestyles. Studying Vibrio cholerae reveals that bacterial populations hedge their bets—some cells disperse while others remain protected. This collective decision-making helps bacteria survive threats and plays a key role in infection and transmission.
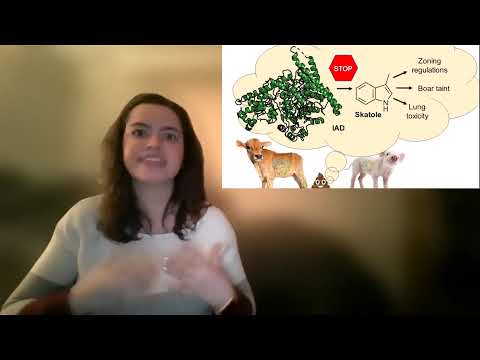
This research identifies and characterizes IAD, a gut-microbial enzyme responsible for producing skatole, a key source of fecal odor. Understanding IAD’s structure and mechanism could help agriculture reduce farm odors, prevent boar taint, and protect cattle health. X-ray crystallography is being used to design inhibitors that block skatole formation.
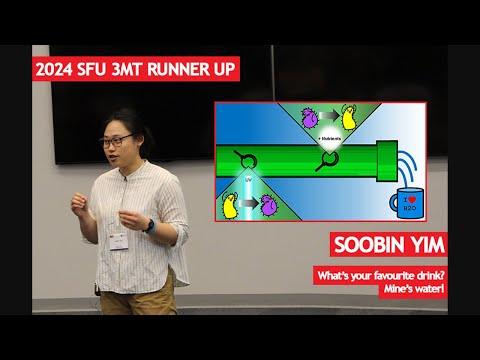
This research examines how microbes in drinking water recover after UV disinfection. By adding nutrients to UV-treated samples and identifying microbes through DNA sequencing, the study tracks which organisms survive, regrow, and thrive over time. The goal is to improve treatment systems and ensure safer, more stable drinking water during distribution.
Pagination
- Page 1
- Next page