This research examines how microbes in drinking water recover after UV disinfection. By adding nutrients to UV-treated samples and identifying microbes through DNA sequencing, the study tracks which organisms survive, regrow, and thrive over time. The goal is to improve treatment systems and ensure safer, more stable drinking water during distribution.
The talk highlights how biology involves unseen interactions and how distinguishing living from dead microorganisms is essential. Using the chemical PMA (propidium monoazide), researchers can identify active pathogens and reduce misinterpretation in diagnostic tests, especially for viruses that cannot be grown in labs. This technique helps improve diagnostics, guide treatments, and advance microbiological research.
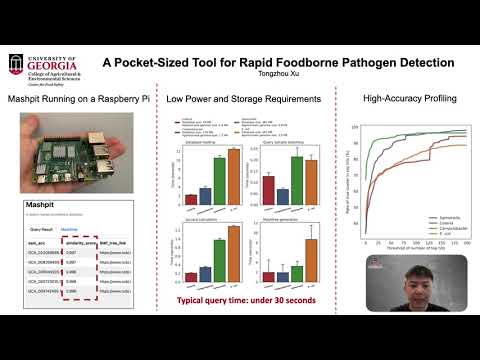
Mashpit is a portable genome-search tool that runs on a Raspberry Pi, enabling rapid, offline screening of Salmonella genomes. Using MinHash sketches, it scans hundreds of thousands of genomes in seconds, offering small or low-resource labs a fast, accessible way to identify related isolates before performing high-resolution follow-up analyses.
My research tackles PFAS (polyfluoroalkyl substances) or “forever chemicals,” found in everyday products and linked to serious health risks. Blood testing shows 95% contamination rates. The project identifies specialised bacteria capable of breaking PFAS down nearly completely within days, offering a promising biological solution to reduce environmental and human exposure to these persistent toxic chemicals.
This research searches for new antibiotics in deep-sea sponge bacteria that have evolved for 580 million years to defend their hosts. By growing these never-before-seen microbes and testing them against superbugs like MRSA, the project aims to discover urgently needed antibiotics to combat rising antimicrobial resistance.
Pagination
- Previous page
- Page 2